Studying Nuclear Dynamics in Living Cells
Live cell imaging has revolutionized our understanding of nuclear organization and dynamics. By visualizing the living nucleus in real-time, we can observe the dynamic properties of the genome and nuclear structures, revealing insights into transcription, replication, DNA repair, and RNA processing that are impossible to obtain from fixed specimens.
Key Advantages:
- • Real-time observation of nuclear processes
- • Quantitative analysis of molecular kinetics
- • Single-cell resolution and heterogeneity
- • Temporal correlation of multiple processes

GFP-tagged PCNA labels subnuclear DNA replication factories
Live Imaging Methodologies
FRAP Analysis
Fluorescence Recovery After Photobleaching measures molecular diffusion and binding kinetics within nuclear compartments.
FCS Techniques
Fluorescence Correlation Spectroscopy analyzes fluctuations in fluorescence intensity to measure molecular concentrations and dynamics.
Time-lapse Microscopy
Long-term imaging of cellular processes to capture nuclear reorganization, cell cycle progression, and stress responses.
Single-Molecule Tracking
Individual molecule visualization to understand local environments and heterogeneity in nuclear processes.
Multicolor Imaging
Simultaneous visualization of multiple nuclear components to study interactions and colocalization.
4D Microscopy
Three-dimensional imaging over time to capture the complete spatial-temporal dynamics of nuclear processes.
Dynamic Nuclear Processes
DNA Replication Dynamics
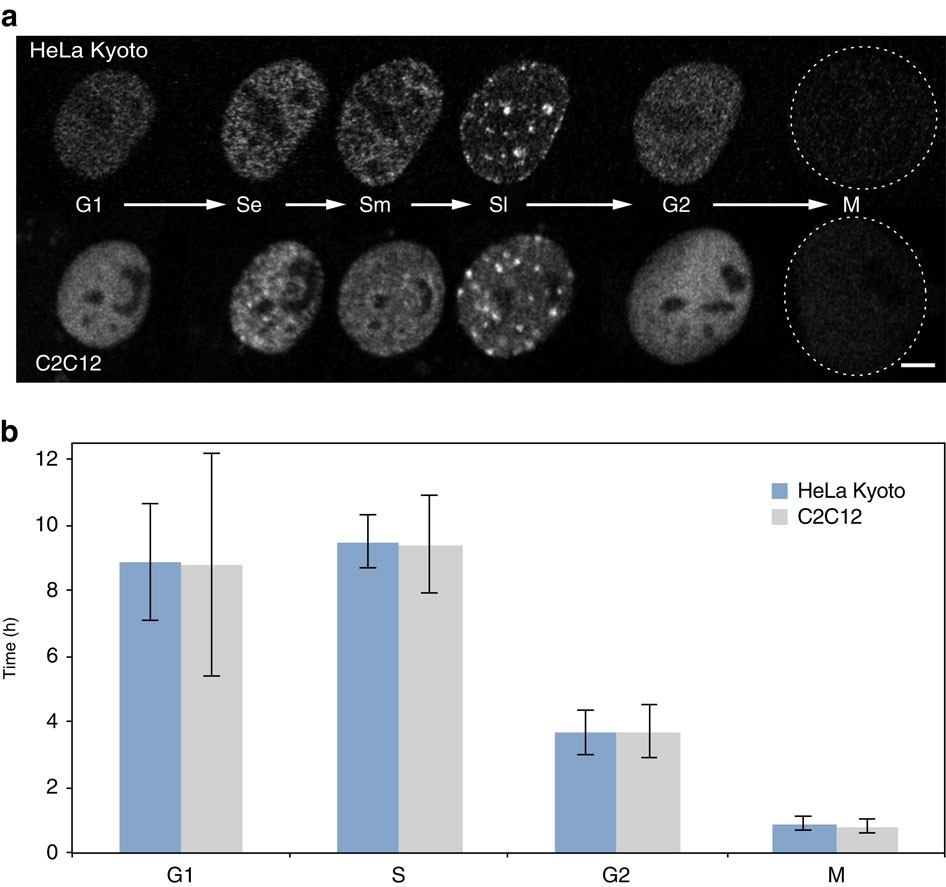
4D visualization of replication foci in mammalian cells
Live imaging reveals the spatiotemporal organization of DNA replication into discrete nuclear factories that change pattern and location throughout S-phase.
Key Observations:
- • Early S-phase: Small, punctate foci (euchromatin)
- • Mid S-phase: Larger foci at nuclear periphery
- • Late S-phase: Intense foci surrounding nucleolus
- • Dynamic protein exchange within stable factories
Transcriptional Dynamics

Localization of replication foci and PCNA in interphase nuclei
Real-time visualization of transcription reveals burst kinetics, polymerase trafficking, and the formation of transcriptional condensates.
Dynamic Features:
- • Transcriptional bursting and stochastic activation
- • RNA polymerase II clustering and tracking
- • Nuclear speckle dynamics and splicing
- • Stress-induced transcriptional reorganization
DNA Repair Focus Dynamics

Evolution of DNA damage-dependent γ-H2AX and 53BP1 focus formation
Live tracking of DNA repair foci reveals the temporal recruitment of repair factors and the coordination of multiple repair pathways.
Repair Dynamics:
- • Hierarchical protein recruitment to damage sites
- • Focus formation and resolution kinetics
- • Checkpoint activation and signaling
- • Repair pathway choice and competition
Nuclear Body Dynamics

Two models explaining nuclear factory dynamics in living cells
Nuclear bodies exhibit complex dynamics including fusion, fission, movement, and compositional changes throughout the cell cycle.
Body Behaviors:
- • Cajal body mobility and gene targeting
- • Nuclear speckle reorganization during stress
- • PML body formation and dissolution
- • Nucleolar dynamics during ribosome biogenesis
Comprehensive Reviews Collection
Explore our extensive collection of comprehensive reviews covering all aspects of nuclear biology and live cell imaging techniques. Each review provides in-depth analysis with the latest research findings.
Live Cell Imaging
DNA Dynamics
Nuclear Bodies
Nuclear Architecture
Biomolecular Condensates
Research Applications
Basic Research
- Nuclear organization mechanisms
- Chromatin dynamics and regulation
- Gene expression control
- DNA repair pathway analysis
Disease Research
- Cancer cell nuclear abnormalities
- Aging and senescence markers
- Neurodegenerative processes
- Viral infection responses
Drug Development
- Target identification and validation
- Compound screening assays
- Mechanism of action studies
- Toxicity and safety assessment
About This Database
This enhanced live cell imaging database integrates cutting-edge research with comprehensive educational resources. All content is based on peer-reviewed research and provides both methodological guidance and theoretical frameworks.
Original content provided by the Michael Hendzel Laboratory, Department of Oncology, University of Alberta, enhanced with comprehensive review collection.
© 2007-2025 Michael J. Hendzel, Ph.D., Department of Oncology, University of Alberta
Enhanced with comprehensive nuclear biology review collection