Chromatin Biophysics and Function: Genome Mechanics
Comprehensive review of chromatin biophysics covering mechanical properties, hierarchical organization, phase separation, and functional integration with quantitative measurements and disease implications
Deep Research Source Review
This page is linked to its matched long-form source review. Read online and submit correction suggestions, or download the full document.
Source file: Chromatin Biophysics and Function Review_.docx | Match confidence: high
Abstract
Chromatin represents one of nature's most sophisticated packaging solutions, compacting approximately two meters of DNA into a nucleus mere microns in diameter. This review explores chromatin as a dynamic, information-rich biopolymer whose physical properties are intrinsically linked to genomic function. We examine the biophysical characteristics—mechanics, electrostatics, and material state—that regulate essential cellular processes from transcription to DNA repair, integrating principles from polymer physics, soft matter science, and molecular biology.
Hierarchical Organization of Chromatin
Structural Levels
- Nucleosome Core: 146-147 bp DNA wrapped around histone octamer
- 10-nm Fiber: "Beads-on-a-string" flexible polymer chain
- Chromatin Loops: TADs with 10-30 minute lifetimes
- Chromosome Territories: Large-scale nuclear organization
Organizational Models
-
Fractal Globule: Compact yet unentangled polymer state allowing dynamic unfolding and refolding
-
Loop Extrusion: SMC complexes (cohesin/condensin) actively extrude loops until encountering CTCF boundaries
-
Chromatin Clutches: Irregularly folded domains with regulated packing density
Chromatin Compaction Levels
Mechanical Properties and Quantitative Measurements
Persistence Length
Naked DNA flexibility
Heterochromatin stiffness
Young's Modulus
Chromatin fiber elasticity
Stiffness Ratio
Heterochromatin vs Euchromatin
Mechanical Properties Comparison
Measurement Techniques
Single-Molecule Methods
- • Optical tweezers for force-extension curves
- • Magnetic tweezers for torsional studies
- • AFM for local stiffness mapping
Ensemble Techniques
- • Microrheology for bulk viscoelasticity
- • SAXS for structural characterization
- • Cryo-EM for native state imaging
Liquid-Liquid Phase Separation and Material States
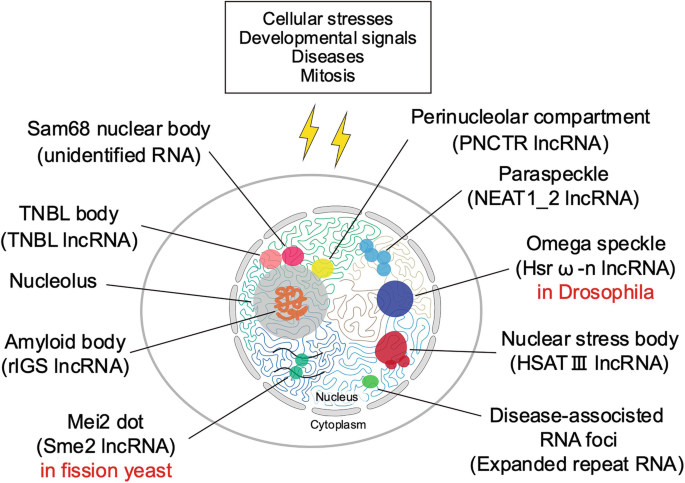
Nuclear Body Formation
Biomolecular condensates form through liquid-liquid phase separation, creating functionally specialized nuclear compartments.
LLPS Mechanisms
Chromatin Material States
Liquid-like
Active euchromatin, rapid exchange
Gel-like
Intermediate compaction states
Solid-like
Heterochromatin, stable domains
Dynamic Properties and Functional Integration
Nucleosome Dynamics
Chromatin Loop Dynamics
Dynamic Timescales
Functional Integration: From Mechanics to Biology
Transcription
Chromatin breathing enables TF access
Replication
Fork progression requires disassembly
DNA Repair
Fluid-like state permits factor access
Recombination
Controlled mobility for locus pairing
Biophysical Regulation Mechanisms
Accessibility Control
- • Chromatin compaction regulates factor binding
- • Nucleosome positioning controls accessibility
- • Phase separation creates functional domains
Search Mechanisms
- • 3D diffusion in nuclear space
- • 1D sliding along DNA
- • Intersegmental transfer between loci
Advanced Research Techniques and Methodologies
3D Structure Mapping
- • Hi-C contact mapping
- • ATAC-seq accessibility
- • ChIP-seq protein binding
- • Super-resolution microscopy
Single-Molecule Analysis
- • Optical/magnetic tweezers
- • FRET dynamics
- • Single-particle tracking
- • Cryo-electron tomography
Computational Modeling
- • Polymer physics simulations
- • Molecular dynamics
- • Coarse-grained models
- • Machine learning integration
Disease Implications and Chromatin Dysfunction
Cancer Biology
- • Altered chromatin compaction in tumors
- • Disrupted phase separation dynamics
- • Oncogene accessibility changes
- • Metastatic chromatin remodeling
Neurodegeneration
- • Protein aggregation in chromatin
- • Loss of nuclear organization
- • Impaired DNA repair mechanisms
- • Aberrant gene expression patterns
Therapeutic Implications
Epigenetic Drugs
Targeting chromatin remodeling complexes, histone modifying enzymes, and DNA methylation machinery to restore normal chromatin states.
Chromatin Engineering
Designing synthetic chromatin architectures and programmable phase separation systems for therapeutic applications.
Conclusions and Future Perspectives
Key Insights
- Chromatin is a dynamic, tunable biomaterial whose physical properties directly regulate genome function
- Phase separation creates functionally specialized nuclear domains with distinct biophysical properties
- Mechanical properties span orders of magnitude and are actively regulated during cellular processes
Future Directions
- Single-cell chromatin biophysics and heterogeneity
- Artificial chromatin systems and synthetic biology
- Therapeutic targeting of chromatin mechanical states
Community Collaboration
This review represents a comprehensive synthesis of current knowledge in chromatin biophysics. We encourage community feedback and collaboration to advance this rapidly evolving field.