Abstract
The mammalian cell nucleus, traditionally viewed as a passive repository for the genome, is now understood to be a dynamic and mechanically responsive organelle. Its biophysical properties are critical for cellular functions ranging from gene regulation and differentiation to migration and division. This comprehensive review analyzes the mechanical nature of the nucleus, synthesizing foundational principles with recent discoveries. We examine the nucleus through its primary mechanical components: the nuclear lamina (acting as a strain-stiffening elastic shell), chromatin (behaving as a viscoelastic polymer gel), and the LINC complex (serving as the crucial bridge transmitting forces from the cytoskeleton to the nuclear interior).
Key Findings
Physical Architecture of the Nucleus
Nuclear Lamina

Strain-stiffening elastic shell composed of intermediate filament proteins (lamins A/C, B1, B2) providing mechanical stability and force transmission.
- • Type A lamins: Lamin A/C
- • Type B lamins: B1, B2
- • Strain-stiffening behavior
- • Disease-associated mutations
Chromatin

Viscoelastic polymer gel with ~2 meters of genomic DNA compacted into ~10 μm diameter nucleus, exhibiting both elastic and time-dependent properties.
- • Heterochromatin (condensed)
- • Euchromatin (relaxed)
- • Viscoelastic properties
- • Gene regulation coupling
LINC Complex
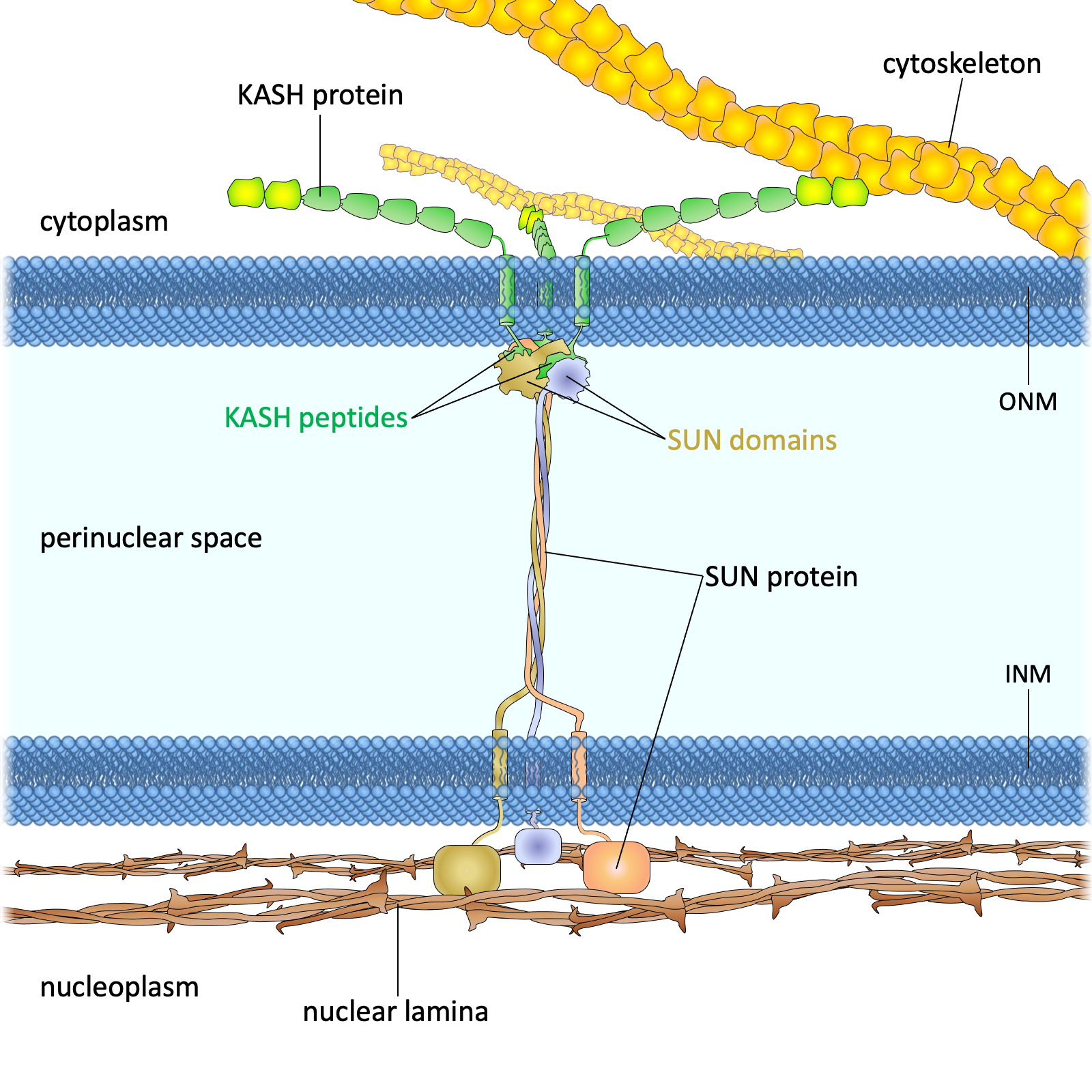
Linker of Nucleoskeleton and Cytoskeleton - SUN/KASH protein bridge transmitting mechanical forces between cytoskeleton and nuclear interior.
- • SUN proteins (inner membrane)
- • KASH proteins (outer membrane)
- • Force transmission
- • Mechanosensing functions
Quantitative Biophysical Measurements
Table 1: Nuclear Elasticity (Young's Modulus) Across Cell Types
| Cell Type | Condition | Young's Modulus (kPa) | Method | Notes |
|---|---|---|---|---|
| hESC | Undifferentiated | 1-2 | Micropipette | Soft, pluripotent state |
| hESC | Differentiated | 6-12 | Micropipette | 6-fold stiffer than undifferentiated |
| hMSC | Undifferentiated | 3.5 | AFM | Mesenchymal stem cells |
| hMSC | Osteogenic | 7.0 | AFM | After differentiation |
| MEF | Wild type | 10.0 | AFM | Mouse embryonic fibroblast |
| MEF | Lmna -/- | 2.5 | AFM | Lamin A/C knockout |
| MCF-10A | Non-tumorigenic | 0.2-0.9 | AFM | Breast epithelial |
| MCF-7 | Cancer | 0.1-0.4 | AFM | Breast cancer - softer |
| HCV29 | Non-tumorigenic | 10-16 | AFM | Bladder epithelial |
| T24 | Cancer | 2.1 | AFM | Bladder cancer - softer |
Table 2: Viscoelastic Parameters of the Nucleus
| Cell Type/Component | Parameter | Value/Range | Method | Significance |
|---|---|---|---|---|
| Neutrophil (whole cell) | Apparent viscosity (η) | 100-200 Pa·s | MA | High viscosity for migration |
| General | Cortical tension | ~30 pN/μm | Various | Surface tension effects |
| Chondrocyte (nucleus) | Instantaneous modulus | 1.8 kPa | AFM | Immediate elastic response |
| Chondrocyte (nucleus) | Equilibrium modulus | 0.5 kPa | AFM | Long-term response |
| Human HSC | Creep exponent (α) | ~0.6 | AFM | Fluid-like behavior |
| Fibroblast | Creep exponent (α) | ~0.2 | AFM | Solid-like behavior |
| MCF-7 | Fast relaxation time | ~0.1 s | AFM | Chromatin/membrane response |
| MCF-7 | Slow relaxation time | ~1.0 s | AFM | Lamina/cytoskeleton response |
| Xenopus nucleolus | Interfacial tension | ~0.4 μN/m | Various | Phase separation dynamics |
| Xenopus nucleolus | Viscosity | 12-32 Pa·s | Various | Internal fluidity |
Table 3: Key Nuclear Proteins, Mechanical Roles, and Associated Pathologies
| Protein/Complex | Mechanical Role | Associated Disease/Pathology | Clinical Impact |
|---|---|---|---|
| Lamin A/C | Primary determinant of nuclear stiffness | EDMD, HGPS, DCM | Muscular dystrophy, premature aging |
| Lamin B1/B2 | Nuclear shape maintenance | Leukodystrophy | Progressive neurodegeneration |
| Emerin | Lamina organization and stability | X-linked EDMD | Cardiac conduction defects |
| LINC Complex | Mechanotransduction, force transmission | Cancer metastasis | Enhanced cell motility |
| Chromatin (general) | Viscoelastic support, gene regulation | Cancer, aging | Altered nuclear deformability |
Data Visualization
Nuclear Stiffness Across Cell Types
Disease vs Normal Cell Stiffness
Experimental Methods for Nuclear Mechanics
Atomic Force Microscopy (AFM)

Other Key Techniques
Micropipette Aspiration
Whole-cell deformation for viscoelastic properties
Optical/Magnetic Tweezers
Precise force application at pN scale
Confined Migration
Microfluidic channels for deformability testing
Brillouin Microscopy
Non-invasive elasticity mapping
RT-DC (Real-time Deformability)
High-throughput mechanical phenotyping
Nuclear Mechanopathology
Laminopathies
Emery-Dreifuss Muscular Dystrophy (EDMD)
- • Lamin A/C or Emerin mutations
- • Reduced nuclear stiffness
- • Cardiac conduction defects
- • Progressive muscle weakness
Hutchinson-Gilford Progeria (HGPS)
- • Lamin A processing defect
- • Abnormal nuclear morphology
- • Premature aging phenotype
- • Cardiovascular complications
Cancer Metastasis
Nuclear Softening
- • Reduced Young's modulus in cancer cells
- • Enhanced deformability for invasion
- • LINC complex alterations
- • Chromatin reorganization
Clinical Examples
- • Breast cancer: 0.1-0.4 kPa (vs 0.2-0.9 normal)
- • Bladder cancer: ~2.1 kPa (vs 10-16 normal)
- • Melanoma: 0.3-0.7 kPa (metastatic variants)
- • Nuclear deformability as biomarker
Functional Integration of Nuclear Mechanics
Development
Plastic-to-stiff transition during stem cell differentiation. ESCs: 1-2 kPa → 6-12 kPa upon lineage commitment.
Mechanotransduction
LINC complex transmits cytoskeletal forces to chromatin, influencing gene expression and cellular responses.
Aging
Nuclear mechanical decline in cellular senescence. Lamin expression changes and chromatin reorganization.
Synthesis and Future Perspectives
Key Insights
- Nuclear mechanics span 3 orders of magnitude (0.1-100 kPa) across different cell types and conditions
- Differentiation consistently increases nuclear stiffness, supporting the mechanostat hypothesis
- Disease states often correlate with altered nuclear mechanics (soft cancers, stiff aged cells)
- Multiple time scales govern nuclear mechanics (ms to hours)
Future Directions
- Single-cell mechanical profiling for disease diagnosis
- Therapeutic targeting of nuclear mechanics in laminopathies
- Real-time mechanomics during cell fate transitions
- Integration with epigenetic and transcriptional networks
Key References & Methods
This review synthesizes data from multiple experimental approaches including atomic force microscopy, micropipette aspiration, optical tweezers, and microfluidic-based assays. Key methodological advances in nuclear mechanics measurement have enabled quantitative characterization across diverse cell types and disease states.
Original Document Source: "Biophysics of the Mammalian Nucleus" - Comprehensive academic review covering nuclear architecture, quantitative measurements, functional integration, and disease implications.